|
We are interested in the methylation of arginine residues of histones and of other chromatin-associated proteins. This posttranslational modification is catalysed by a family of enzymes, the PRMTs (protein arginine methyltransferases), which are evolutionary conserved in eukaryotes from fungi to plants and mammals (Bedford and Clarke, 2009). In humans the enzyme family consists of eleven members (PRMT1-PRMT11). PRMTs share a central highly conserved catalytic domain and perform mono- and dimethylation of the guanidino group of the arginines using S-adenosylmethionine (SAM) as methyl donor.
PRMTs are implicated in the regulation of a plethora of cellular processes: RNA processing, ribosome biogenesis, nucleo-cytoplasmic transport and chromatin-associated processes, such as DNA-repair, imprinting and transcriptional regulation (Bedford and Clarke, 2009). Knockout mice revealed an essential function of PRMT1 and PRMT4 during embryogenesis (Pawlak et al., 2000; Yadav et al., 2003). Furthermore, deregulated expression of PRMT1, PRMT4 and PRMT5 was found to be associated with the occurrence of certain tumor types, for example PRMT4 is upregulated in breast cancer with poor prognosis (El Messaoudi et al., 2006; Majumder et al., 2006; Pal et al., 2007).
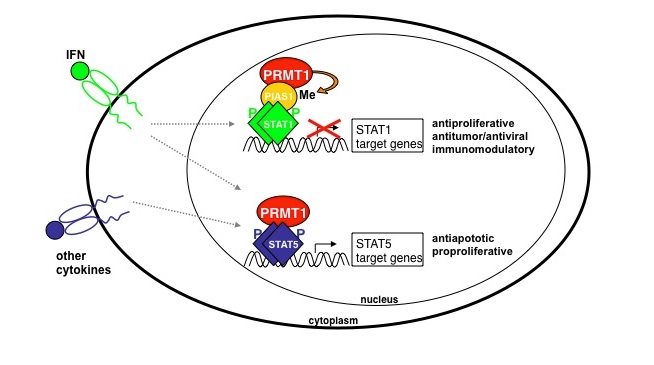
Our work concentrates on the chromatin-regulatory functions of arginine methylation, To fulfil their function as transcriptional coregulators the enzymes interact with distinct transcription factors and are recruited to the corresponding target gene promoters. PRMTs affect gene activity either positively or negatively in a methyltransferase-dependent manner by modifying chromatin-associated proteins. Using microarray-based expression profiling, we found that the transcription factor STAT5 is coactivated by the histone methyltransferase activities of PRMT1 and PRMT4 (Kleinschmidt et al., 2008). Likewise, other reports suggested that arginine methylation by PRMT1 regulates cytokine, in particular IFN, signaling. PRMT1 facilitates the expression of cytokine genes by coactivating the pro-inflammatory transcription factor NFAT (Mowen et al., 2004). On the other hand, PRMT1 interacts with the cytoplasmic domain of the interferon α (IFNα) receptor and depletion of PRMT1 alters IFNα-induced growth arrest (Abramovich et al., 1997; Altschuler et al., 1999). Recently, we could show that PRMT1 methylates PIAS1, a nuclear inhibitor of the transcription factor STAT1, and this modification triggers interaction between STAT1 and PIAS1 resulting in the shut-down of STAT1-mediated gene expression (Komyod et al., 2005; Weber et al., 2009).
In summary, our data (summarized in the figure) suggest that PRMT1 on the one hand inhibits the transactivation function of STAT1 and on the other hand is a coactivator of STAT5-mediated transcription. This is interesting as deregulated STAT activity correlates with the onset of leukemia and melanoma, e.g. in melanoma cells STAT5 overexpression antagonizes the IFNγ-inducible anti-proliferative effect of STAT1 (Wellbrock et al., 2005). Therefore, PRMT1 is potentially very useful target in therapies of deregulated STAT signal transduction and inhibition of PRMT1 might be beneficial.
In this project we have the following goals:
(1) to identify the mechanism of the corepressor function of PRMT1 in STAT1 signaling
(2) to analyze the genome-wide target gene expression (in particular with respect to STAT1 and STAT5 target genes) and functional consequence of PRMT1-depletion in IFN-resistent melanoma cells and
(3) to investigate the function of PRMT1 in transcriptional regulation by NF-kB, another transcription factor relevant for inflammation and cancer.
References:
Abramovich, C., Yakobson, B., Chebath, J. and Revel, M. (1997) A protein-arginine methyltransferase binds to the intracytoplasmic domain of the IFNAR1 chain in the type I interferon receptor. Embo J, 16, 260-266.
Altschuler, L., Wook, J.O., Gurari, D., Chebath, J. and Revel, M. (1999) Involvement of receptor-bound protein methyltransferase PRMT1 in antiviral and antiproliferative effects of type I interferons. J Interferon Cytokine Res, 19, 189-195.
Bedford, M.T. and Clarke, S.G. (2009) Protein arginine methylation in mammals: who, what, and why. Mol Cell, 33, 1-13.
El Messaoudi, S., Fabbrizio, E., Rodriguez, C., Chuchana, P., Fauquier, L., Cheng, D., Theillet, C., Vandel, L., Bedford, M.T. and Sardet, C. (2006) Coactivator-associated arginine methyltransferase 1 (CARM1) is a positive regulator of the Cyclin E1 gene. Proc Natl Acad Sci U S A, 103, 13351-13356.
Kleinschmidt, M.A., Streubel, G., Samans, B., Krause, M. and Bauer, U.M. (2008) The protein arginine methyltransferases CARM1 and PRMT1 cooperate in gene regulation. Nucleic Acids Res.
Komyod, W., Bauer, U.M., Heinrich, P.C., Haan, S. and Behrmann, I. (2005) Are STATS arginine-methylated? J Biol Chem, 280, 21700-21705.
Majumder, S., Liu, Y., Ford, O.H., 3rd, Mohler, J.L. and Whang, Y.E. (2006) Involvement of arginine methyltransferase CARM1 in androgen receptor function and prostate cancer cell viability. Prostate, 66, 1292-1301.
Mowen, K.A., Schurter, B.T., Fathman, J.W., David, M. and Glimcher, L.H. (2004) Arginine methylation of NIP45 modulates cytokine gene expression in effector T lymphocytes. Mol Cell, 15, 559-571.
Pal, S., Baiocchi, R.A., Byrd, J.C., Grever, M.R., Jacob, S.T. and Sif, S. (2007) Low levels of miR-92b/96 induce PRMT5 translation and H3R8/H4R3 methylation in mantle cell lymphoma. Embo J, 26, 3558-3569.
Pawlak, M.R., Scherer, C.A., Chen, J., Roshon, M.J. and Ruley, H.E. (2000) Arginine N-methyltransferase 1 is required for early postimplantation mouse development, but cells deficient in the enzyme are viable. Mol Cell Biol, 20, 4859-4869.
Wellbrock, C., Weisser, C., Hassel, J.C., Fischer, P., Becker, J., Vetter, C.S., Behrmann, I., Kortylewski, M., Heinrich, P.C. and Schartl, M. (2005) STAT5 contributes to interferon resistance of melanoma cells. Curr Biol, 15, 1629-1639.
Yadav, N., Lee, J., Kim, J., Shen, J., Hu, M.C., Aldaz, C.M. and Bedford, M.T. (2003) Specific protein methylation defects and gene expression perturbations in coactivator-associated arginine methyltransferase 1-deficient mice. Proc Natl Acad Sci U S A, 100, 6464-6468.
|