|
B4: SUMO-dependent transrepression of inflammation induced gene expression |
|
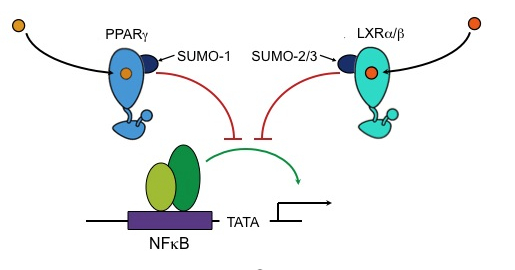
Transrepression of NFκB-activated proinflammatory genes by peroxysome proliferator-activated receptor-γ (PPARγ) and liver X receptor (LXR) is dependent on SUMOylation. The molecular mechanism by which SUMO-modification of these nuclear receptors effects transrepression is not fully understood.
By using a genome-wide RNA interference screen, we have recently identified SUMO-dependent repression components (1), and described SUMO-modification of transcription factors as a novel mechanism for the initiation and formation of repressive local heterochromatic-like states (2). Guided by the mechanistic clues provided by these findings, we hypothesize that similar chromatin changes could take place upon transrepression of proinflammatory genes by SUMOylated PPARγ and LXR. To address this question, we will perform RNAi and chromatin immuonprecipitation experiments.
PPARγ und LXR are modified by different SUMO paralogs (PPARγ by SUMO1 and LXR by SUMO2/3). Hence, it is very likely that different gene and signal-specific transrepression programs are affected by SUMO1 and SUMO2/3. To identify proflammatory genes that are specifically affected by different SUMO paralogs we will perform RNAi experiments in combination with microarray analyses.
References:
(1) Stielow B, Sapetschnig A, Krüger I, Kunert N, Brehm A, Boutros M., and Suske G (2008) Identification of SUMO-dependent chromatin-associated transcriptional repression components by a genome-wide RNA interference screen. Mol. Cell 29, 742-754.
(2) Stielow B, Sapetschnig A, Wink C., Krüger, I., and Suske G. (2008) SUMO-modified Sp3 represses transcription by provoking local heterochromatic gene silencing. EMBO Rep. 9, 899-906.
|
|
Last Updated on Monday, 15 June 2009 08:24 |