
Z1: Genomics Unit


Prof. Dr. Thorsten Stiewe
Prof. Dr. Rolf Müller
Institut für Molekularbiologie und Tumorforschung
Phone: +49 6421 2863710
Fax: +49 6421 2865010
Phone: +49 6421 2863710
Fax: +49 6421 2865010

| Z1: Genomics Unit |
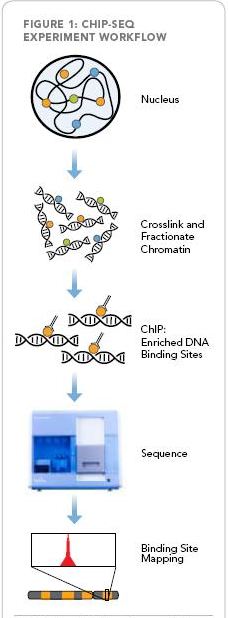 A Genomic and Microarray Unit was installed in 2001 at the Institute of Molecular Biology and Tumor Research (IMT) at the Philipps University Marburg to provide researchers with state-of-the-art methodology and to further boost microarray technology. The unit provides cDNA microarrays, miRNA-arrays, high throughput RNAi screenig of shRNA-librarys and screening of cells by laser scanning microscopy. Within the LOEWE research cluster a new technology, ChIP-Seq, will be established as a key technology for the genome-wide detection of transcription and chromatin regulator binding sites. A Genomic and Microarray Unit was installed in 2001 at the Institute of Molecular Biology and Tumor Research (IMT) at the Philipps University Marburg to provide researchers with state-of-the-art methodology and to further boost microarray technology. The unit provides cDNA microarrays, miRNA-arrays, high throughput RNAi screenig of shRNA-librarys and screening of cells by laser scanning microscopy. Within the LOEWE research cluster a new technology, ChIP-Seq, will be established as a key technology for the genome-wide detection of transcription and chromatin regulator binding sites.For conventional ChIP-on-chip-technology immunoprecipitated DNA fragments are amplified, fluorescence labeld and hybridized to oligonucleotides immobilized to high density microarrays, thus allowing for the detection of transcription factor binding sites. Due to array space limitations the ChIP-on-chip-technology is, however, limited to hybridization of pre-selected oligonucleotides making it impossible to screen transcription factor binding sites on a genome-wide level. The ChIP-Seq technology has significantly advanced the genome-wide detection approach for transcriptional binding sites. By this new technique, the direct sequencing of precipitated DNA is possible, thereby bypassing the spatial limitations of the ChIP-on-chip-technology and allowing for the detection of all factor binding sites at a genome-wide level. This new powerfull, ChIP-Seq technology will be implemented within the established Microarray Unit at the IMT and will be made available to the LOEWE scientists during the course of 2010.
Fig. 1: Protein-chromatin interactions are first crosslinked in situ. Specific DNA fragments are co-immunoprecipitated and sequenced to identify genome-wide sites associated with a factor or modification of interest (see also homepage of Illumina) |
| Last Updated on Thursday, 17 February 2011 12:42 |

